Personal
Career summary (see CV for more information)
I retired from the full time Yale faculty in July 2021 and moved to Berkeley, CA, where I continue to do research at home and in the laboratory of David Drubin at UC Berkeley. Bill Earnshaw and I have authored of four editions of the textbook “Cell Biology” with illustrations by Graham Johnson.
In addition to research and teaching, I chaired my departments at the Johns Hopkins Medical School and Yale University, served as President of the Salk Institute for Biological Studies and was Dean of the Graduate School of Arts and Sciences at Yale. I served as president of both the American Society for Cell Biology and the Biophysical Society and in leadership positions at the National Academy of Sciences. For more details, see an autobiography at https://doi.org/10.1146/annurev-cellbio-100818-125427.
My honors include the Gairdner International Award, E.B. Wilson Medal from the American Society for Cell Biology, the Medal of Science from the State of Connecticut and election to the American Academy of Arts and Sciences, National Academy of Sciences and National Academy of Medicine.
Research summary (for more information see page on Research)
Starting in the 1960s, I have investigated along with members of my laboratory the molecular basis of cellular movements and cytokinesis using a combination of biochemistry, biophysics, microscopy and computational modeling.
Actin-based cellular movements are essential for shaping organs during embryonic development, defense against microorganisms and wiring the nervous system. Movement of cells out of primary tumors is the chief cause of mortality in cancer. Cytokinesis is essential for the replication of all cells and proliferation of cancer cells.
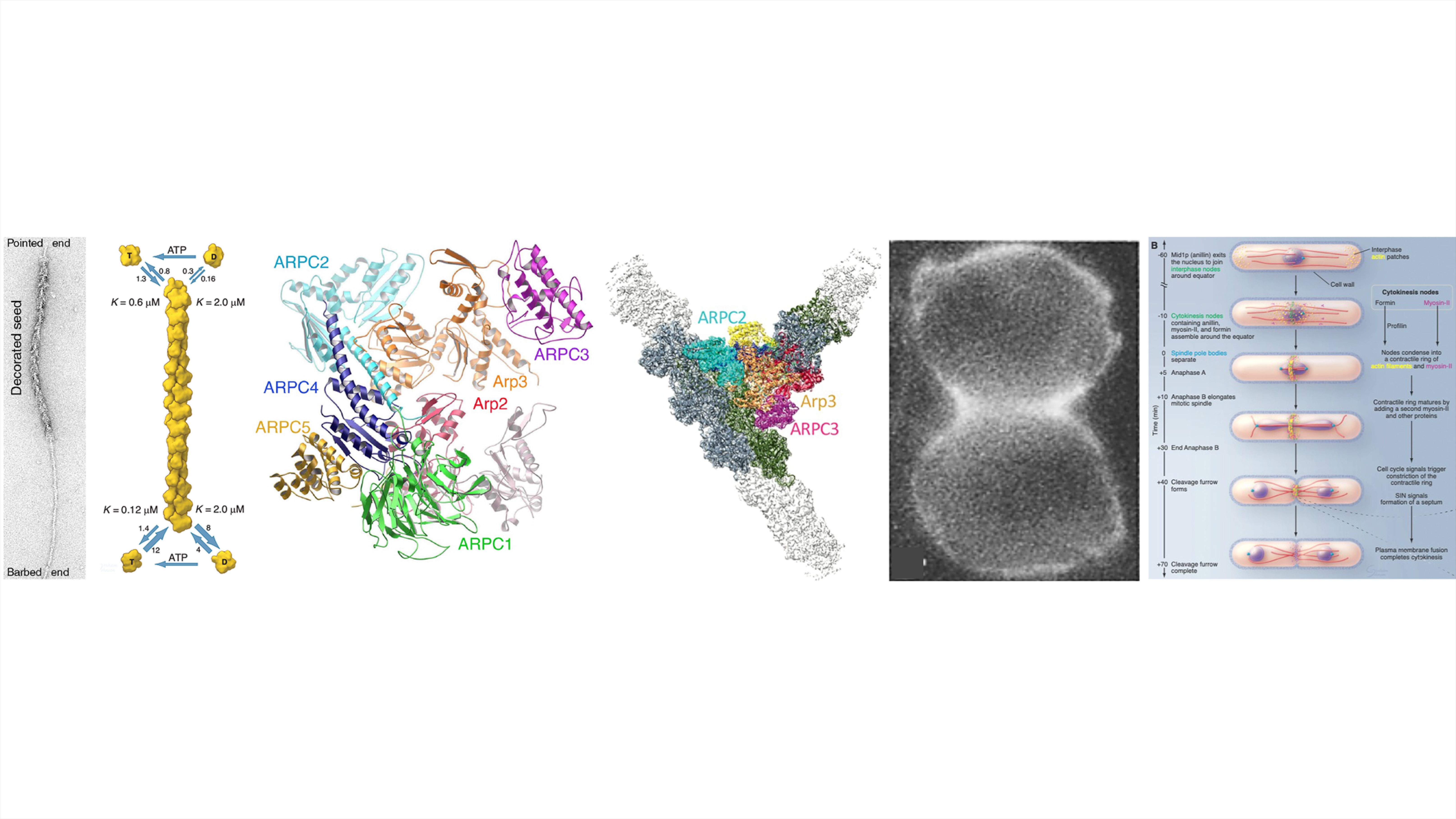
My laboratory discovered and characterized many proteins that produce forces for cells to move including the first unconventional myosin (myosin-I), Arp2/3 complex and capping protein, all originally isolated from Acanthamoeba. We combined microscopy, biochemistry, biophysics, molecular genetics and mathematical modeling to provide the quantitative evidence required to formulate and test a detailed molecular explanation for how Arp2/3 complex stimulates the assembly of branched actin filaments that produce forces for cellular movements and endocytosis. We combined quantitative measurements of the time course of the appearance and disappearance of the participating proteins at site of endocytosis in fission yeast with mathematical modeling to confirm the molecular details and physics of the force-producing process.
Postdoc Keigi Fujiwara discovered myosin-II in the cleavage furrow of Hela cells in 1976, initiating our research on cytokinesis. In the late 1990’s the lab switched to using fission yeast to investigate cytokinesis. We characterized the participating proteins (actin, two isoforms of myosin-II, formin Cdc12, IQGAP Rng2, anillin Mid1p, F-BAR Cdc15, profilin, cofilin and capping protein) and measured the numbers of these proteins over time in the cytokinesis structures of live cells. This work culminated in the first molecularly explicit mathematical models and computer simulations of the mechanisms that assemble and constrict the cytokinetic contractile ring.